Being Oncomine branded, can you do lot matching?
You should expect each order of a panel to be a new lot number. However, there is a possibility that 2 orders provided on the same SO could be the same lot number.
I see some tumor types are missing, will these get added eventually, like brain or sarcoma?
We are investigating the addition of new tumor types and new genes in the future. Please contact support or provide assay feedback on AmpliSeq.com for any additions you’d like to see, including additional genes. Specific design feedback should be provided using the Give Solution Feedback option in the More Actions drop-down menu on the panel design page. To provide general feedback or request help, use the Assay Design Support link in the footer of each page on AmpliSeq.com.
What is the difference between a core panel, predesigned panel, and customized panel?
All genes in stock have been tested together to assure they perform well together in any permutation. The core panels and the genes in these panels have been tested to the highest performance compared to other genes in inventory. The core panels also include corresponding analysis workflows in Ion Reporter software. Customized panels do not include default analysis workflows and will require a copy-edit of one of the default workflows. Predesigned panels are similar custom panels and do not include a default analysis workflow in Ion Reporter software, but can be ordered without panel finalization like the core panels.
What is the maximum and minimum number of genes/amplicons allowed?
The minimum number of amplicons is 24 (12 per pool). The maximum number of amplicons is 5000 or 150 genes, whichever is reached first.
The added library kits have different product numbers than I’m used to. Are they different than the ones available on Thermofisher.com?
The library kits are the same as those commercially available, simply under a different product number. These library kits are only available with the Oncomine tumor specific panels from AmpliSeq.com.
What rationale can you provide for the core panel gene content? What are associated research genes?
Core panel content is designed to be most relevant for clinical research of the tumor type based on drug labels, guidelines and clinical trials. The panel can be customized if you prefer to add or replace certain genes.
Associated research genes may also be of interest based on our curation of the relevant literature.
Why can’t I add RNA content to a new panel created from a gene list?
Only panels derived from an existing Oncomine tumor specific panels are currently supported for having the RNA content.
How can I find all my custom panels that contain fusion variant support?
We have added a filter category to your “My Designs” section, under the Oncomine Tumor Specific tab, to include designs that exclusively contain DNA and Fusion support.
When I create a custom design for an Oncomine tumor specific panel that supports fusions, how do I ensure that the fusion panel is included or removed?
During the finalization process, the user will be given the option to select whether the design should be finalized as a DNA only design or a DNA and Fusions design. Making the DNA and Fusions design selection will ensure that the Oncomine solid tumor fusion panel is included, and the design categorize in the right section under My Designs.
What if I just want amplicons to cover hotspots for a certain gene, not the full gene design?
Each gene is available as is without modification. Please contact support for help designing amplicons covering only your genomic location of interest. An assay feedback tool is available on the designer – please submit any requests for modification to a gene design.
The amplicon count for the DNA pools of my Oncomine tumor specific panels in the design page is different than the amplicon count on the tube label. Is this an error?
No, this is not an error and is due to the presence of sample ID amplicons in the DNA pools, a design feature in our Oncomine tumor specific panels. Our user interface shows the number of TARGET amplicons and does not include sample ID amplicons in the panel page display or order files. Consequently, you will see a lower amplicon count in the user interface (panel page) compared to the DNA pool tube labels, which are instead configured to report the total number of amplicons present in the tubes. The correct number of target amplicons and of total amplicons is therefore shown both on the AmpliSeq.com user interface and on the DNA pool tube labels, respectively, for your Oncomine tumor specific panels.
Why can I only order 500 genes in my panel?
For this version of the software, we’ve set an ordering limit to 500 genes or 15,000 amplicons per panel due to manufacturing restrictions.
Why is there a limit on the number of genes that I can add to my panel?
Since the order limit is set at 500 genes per panel, it becomes impractical to allow a large number of genes into the Grid or Table view, which will need to be deselected in order to make the design orderable. For this reason, we’ve introduced a limit on the number of genes that can be added to an On-Demand panel.
Can I edit the content once I’ve created a design?
Yes, an On-Demand design can be edited after it has been created as long as it has not been locked by one of the following actions:
- The panel is finalized by the user by clicking the “Finalize Design” button in the panel page.
- A spike-in panel has been created for the panel.
This is different from Made-To-Order designs, which can only be edited in the “Draft” mode and become locked once the job has been submitted and the Results reported.
By finalizing an On-Demand design, the user is agreeing to locking the design from further editing. Once done, user will be able to download panel files (including target bed file), review panel design information, and place an order/request a quote for the panel, but not to edit the design content.
Note: only checked genes will be submitted to the finalized design.
The content of a locked On-Demand panel may be edited creating a panel clone by clicking on the Clone Panel button. A new unlocked panel with the same content, but under a new IAD code, will be automatically created.
Can I download my list of targets once I’ve created a design?
Yes, select the “Export targets” button to download the list as a CSV file. This will export all the selected targets displayed in the user interface.
Once I have created a design, can I add more content from a Disease Research Area (DRA)?
Not directly. Currently, we do not allow the addition of DRA content or hierarchy levels to an existing design, only when the design is being created. The solution is to create a new design with the desired DRA content or hierarchy levels and add content from inventory as desired when in the unlocked design state.
-
When you click the “Order” button, only genes that are available as On-Demand genes, and which you selected (green), will be ordered. If you create a Spike-in panel, that panel needs to be submitted and ordered separately by visiting the results page of that panel. The Spike-In panel is processed as a made-to-order custom panel and follows the same design submission and ordering process, including separate manufacturing and shipping.
Can I edit my design once I’ve placed an order?
No, once you’ve placed an order, the design cannot be edited because the necessary files needed for analysis by Torrent Suite and Ion Reporter Software need to remain in sync with the material you ordered. If you need to edit your design, select the “Clone” option to copy the panel design. A new IAD number will then be assigned to your design, and you will have the option to edit the design content. Cloning a panel will copy the entire design, selecting any available genes from inventory including any from the Spike-In panel that is now available from potential inventory updates.
Can I reorder a design once I’ve placed an initial order?
Yes, you can always go back to your ordered design and place a new order. If you want to order multiple copies of a panel, the solution is to either order a larger reaction size of the panel (ie one 96 rxn vs three 24rxn) or to go back to your design and add to cart again.
What is the annotation source and version that is used to recognize gene symbols when creating an On-Demand Panel?
The source of annotations is RefGene and the version that we’re using is version v74.
Are untranslated regions (UTRs) included with a gene design?
No, only the coding DNA sequence (CDS) region of a gene is included as part of an On-Demand gene design. If UTRs or amplicons are desired, please contact the support team for potential spike-in solutions via Made-to-Order pipeline.
Are UTR-only genes supported? What about pseudogenes?
No, only genes containing CDS regions are supported. At this time, pseudogenes are not supported.
What is the padding used for gene designs?
The padding for every On-Demand gene design is either 5 bp or 25 bp on the 5’ and 3’ ends.
Can I share my design with a collaborator the same way I do with a Made-to-Order (ie custom) design?
Yes, we do support sharing. Alternatively, you can also export the list of targets, and share that list with your collaborator. The design they create will be identical to yours if the list of targets is the same.
What does the “Score” mean?
The “Score” ranks the relationship between a gene and a disease and it is used to associate genes to disease categories in the Disease Research Areas tool. It takes into account both the strength and number of gene-disease pairs. The algorithm to determine scoring is proprietary. Once genes are added to an AmpliSeq On-Demand panel, they are sorted by the value of this score by default. If needed, switching to Table view on the panel page allows to visualize the Score value for each gene in the panel.
What is “in-silico” coverage?
“In-silico” coverage is defined by the percentage of bases that are covered by the tiling of amplicons. This number is a computer-based calculation and should not be confused with experimental coverage, which represents the actual performance of the panel in the lab. We have wet-lab tested all inventoried content in-house.
-
The number of reads spanning is counted for each base across all padded coding exons of a gene. An average value is calculated for all the bases, and the percentage of bases with read counts above 20% of the average value is defined as “Gene Uniformity”.
Have you tested all possible gene combinations for primer-primer interactions?
No, the number of possible combinations is astronomical and it is not possible to test for all possible combinations in the lab. What our in-house R&D team has done is use computer-based searches to reduce as much as possible the occurrence of primer-primer interactions. The risk is not negligible but deemed very low, backed further by the number of satisfied customers. We have observed << 1% amplicon drop-out due to suspected primer-primer interactions.
Further, we cannot guarantee specifications regarding off-target. We support the in-house GBU and coverage indicated in the IGV viewer and will do our best help troubleshoot if there are any issues that we believe are due to the design or manufacture or our panels.
If my design has a banner indicating a gene(s) has amplicons greater than designed 325bp, what should I do?
The default sequencing protocol for On Demand panels is 200bp and 550 nucleotide flows on any chip. If the user wants end to end reads on amplicons greater than 325bp (which can increase detection and accuracy of variant calls by reading both strands), we recommend increasing the number of flows to 650 and using 510, 520 or 530 chips. Amplicons between 275bp and 325bp can use the default workflow. Note, with >550 flows, only one sequencing run per S5 initialization will be possible. The user can determine the size of the insert from the bed file and then identify the amplicons greater than 325bp by adding on the length of the adapters/barcodes (~48bp).
Refer to the Ion AmpliSeq Library Kit Plus User Guide (MAN0017003) for more information.
What is the shelf life for these panels? How should they be shipped/stored? Do you supply a COA?
Shelf life: 730 days based on the gene with the earliest manufacturing date.
Shipped: Room Temperature.
Storage: -20° C for longest stability. Avoid freeze-thaw cycles.
COA: As this is a custom product, we can only provide a Certificate of Conformance (COC) upon request, as opposed to off-the-shelf products for which a COA is available for download.
What is the turnaround for these panels once I place an order?
As these are custom panels, made upon receipt of the order, the turnaround times can vary depending on geographical location but we aim for 2-3 weeks. We have no guarantees for TAT.
How do I scale up my panel?
We currently do not offer On-Demand panels in larger reactions packs than 32 rxn for Chef or 96 rxn for manual.
I recently received an AmpliSeq On-Demand panel where the number of amplicons per pool on tube labels is higher than that reported on the panel page and in designed.bed file. Is this expected?
In order to obtain proper performance, some amplicons are added multiple times into their respective pool. This is by design. AmpliSeq on-Demand panels that include such amplicons will have this discrepancy. This is not an issue. The panel content is as expected, and the functional number of amplicons is the one displayed on panel page and designed.bed file.
When should I use Ion AmpliSeq HD (ASHD) and not “regular” Ion AmpliSeq (AS)?
It depends on the application. If you have an application that requires ultra-high sensitivity, then ASHD is recommended. If you have applications that require high multiplexing, then regular AS is the preferred option.
What is the maximum number of amplicons per pool that are supported by ASHD?
ASHD currently supports up to 4999 amplicons (9999 oligos) per design. This limit is enforced in AmpliSeq.com.
How many reactions worth of material can I expect in my order of an ASHD panel?
The approximate number of reactions worth of material provided for ASHD panels is 3,000 reactions for 1-pool panels and 6,000 reactions for 2-pool panels (manual library preparation).
Designs are only available for cfDNA and FFPE, what about Germline applications?
Germline applications may be considered in the future, depending on the relevance use of ASHD technology.
What are the variant types supported by ASHD?
The currently supported variant types are, SNVs, small Indels, Fusions from a pre-designed list, and CNV through design best practices available in AmpliSeq.com.
What happens if my desired fusion is not available in AmpliSeq.com?
In this case, a request to the AmpliSeq Custom Services team will need to be submitted. Contact your local sales or support representative to learn more.
Will the AmpliSeq Custom Services team support specialty designs for ASHD?
Yes, the AmpliSeq Custom Services team will be supporting specialty designs for ASHD. Submission of specialty ASHD designs to the AmpliSeq Custom Services team should follow the same path for AS design requests, which involves contacting your sales or field support contacts for help.
Only gene expression controls are available when creating a fusion design, what about custom designs for gene expression assays, are these supported by ASHD?
Automated gene expression designs are not currently supported in AmpliSeq.com. However, custom gene expression designs can be created by the AmpliSeq Custom Services team. Contact your local sales or support representative to learn more.
The new Oncomine cf PanCancer panel contains DNA and RNA targets in 1-pool, can you create a similar design with ASHD?
Yes, but only through our AmpliSeq Custom Services team. Designs created in AmpliSeq.com are limited to 1-pool for DNA hotspots, 1-pool for RNA fusions, and 2-pools for DNA gene designs.
Why are the FWD and REV primers kept in separate pools?
The primers used in ASHD are more complex than for regular AS, so it has been recommended to keep them in separate pools for storage and long term stability. FWD and REV primers should only be mixed at the time the libraries are created. Please refer to ASHD Library Kit User Guide for more information.
Does the chip selection influence the LOD that can be achieved with AmpliSeq HD?
Yes, the chip selection is governed by the following two considerations:
Chip throughput requirement to achieve the desired LOD and the number of samples per chip for the specific assay panel with the specific number of amplicons. Refer to Chapter 6 of the User Guide (MAN0017392) to learn more about the approximate number of libraries supported per chip type based on the number of amplicons per library. See Appendix C to learn about the coverage read requirements based on input amount and target LOD.
Lengths of amplicon for AmpliSeq HD designs supported by different chips. For cfDNA/FFPE designs with amplicon size at 75-125bp, your assay is supported on the 530, 540 and 550 chips. However, for the FFPE only design with amplicon size at 125-175bp, your assay is officially supported by only the 530 chip. If higher throughput is needed, 540 can be used but may yield lower number of aligned reads (~50 million reads) on a 540 chip. We do not recommend the use of the 550 chip for the FFPE only design.
Why can I only use the 530 chip for FFPE designs with amplicon size of 125-175bp?
Our studies have shown that as the chip capacity increases, the end-to-end read percentage decreases. Since AmpliSeq HD requires that both barcodes (5’ and the 3’ end) be read correctly, reads that do not have good end-to-end get removed, and hence the total number of families gets lowered and we’re unable to achieve lower limits of detection.
What does “dual use cfDNA/FFPE design” mean?
Dual use designs are designs that can be used for multiple DNA types such as DNA extracted from cell free or FFPE stored tissue. In the case of cfDNA/FFPE, small sized designs (75-125bp) have been shown to be suitable for both types of DNA.
Can I use a 520 chip with AmpliSeq HD?
Yes, 520 chips may be used, but they required custom Torrent Suite parameters. Default parameters are only provided for the 530 chip; consult your local Field Bioinformatics Scientists for help with setting up the parameters in Torrent Suite for the 520 chip.
-
AmpliSeq HD panel with greater than 3 pools can be occasionally created in copy amplicon designs (Subset design), in order to resolve primer conflicts. However these solutions cannot be manufactured, hence the Preview order button is disabled. For support on possible primer conflict resolution strategies, please contact support Americas, EMEA, Greater China, South Asia, Japan for assistance.
-
Refer to the link for a table describing the number of reactions that you can expect, depending on the size of your panel.
-
Refer to the link for a table describing the number of reactions that you can expect, depending on the size of your panel.
-
Refer to the link for a table describing the number of reactions that you can expect, depending on the size of your panel.
-
No, not currently. The only format available for RNA designs is in Tubes Only format.
Why do we provide pooled and plated primers?
To maximize convenience and flexibility. Pooled primers can be used immediately. Plated oligos can be used to: 1) Rebuild the same pool, 2) Rebuild a pool with fewer primers.
Can I add a few more genes to a set of previously ordered primer?
Currently, the only way to do this is to duplicate the target list in a new version, add the new genes and resubmit. As long as the same design attributes are set (CDS/all exons 150/200) as previously used, the genes from the previous set will have the same design.
I am not in the United States. Will my order be shipped directly to my lab, or first to a local Thermo Fisher distribution center, then to my lab?
This depends. With the exception of orders from Europe that are processed in the U.K., all other international orders are first processed by the North America customer service team, who sends the form to the local customer service team for verification and final changes. The local customer service team works with the customer to determine the best route for a shipment, and the decision is made by the local customer service team. Shipment to a distribution center is slower, but it is significantly less expensive for Thermo Fisher, and could potentially result in fewer customs issues and tax charges. Direct shipment is generally faster, but this adds additional shipping costs for Thermo Fisher, and there may be customs and/or tax implications.
If I have a few regular primers for a region and I know they are working, can I add these primers to my AmpliSeq design?
Yes, you will need to use the "Copy Amplicons" function to ensure the amplicons used in the original design are used for your new design.
Can I add primers manually, afterwards, to completely cover a region?
Yes, you can add amplicons to your assay using the process known as "Spike-In". Check with your local Field Support team for guidance on how to add the new targets, and make sure you make the selection for 50X primer concentration at the time of ordering.
Is there a minimum order for AmpliSeq?
Ion AmpliSeq Custom Panels range from 12 amplicons* to 6,144 amplicons per tube. The minimum of 12 amplicons is due to chemistry performance limitations with low number of amplicons per pool. The exception to this limitation are spike-in designs which allows < 12 amplicons/pool, since these panels are exclusively meant to be used to physically add amplicons to larger main designs to expand their content. Note: if an MTO DNA design panel has fewer than 48 amplicons it is subjected to our minimum order quantity policy of 48 amplicons (96 oligos) and its associated pricing.
*Target regions can be as small as 1 kb and can go up to 5 Mb.
-
Both. Each custom primer pool is delivered as both a pre-pooled tube and as individual primer pairs plated into 384-well plates. Small orders of up to 96 amplicons per pool will contain 750 pre-pooled reactions and individual primer pairs sufficient for 1,500 reactions. Larger orders of more than 96 amplicons per pool will contain 3,000 pre-pooled reactions and individual primer pairs sufficient for 6,000 reactions.
-
Email us at genomicorders@thermofisher.com
or call 1-800-955-6288, x46636. Please use your Ion AmpliSeq Design ID number when referring to your order. Please contact your local customer service outside of North America.
What is the Custom oligo cancellation policy?
There is no guarantee of cancellation of a custom oligo order. Please contact your local customer service representative for more information and options. On occasion, customer service can intercept an order and is able to cancel it prior to synthesis. You must call ASAP: 1-800-955-6288, x46636.
-
Refer to the link for a table describing the number of reactions that you can expect, depending on the size of your panel.
-
Refer to the link for a table describing the number of reactions that you can expect, depending on the size of your panel.
AmpliSeq Made-to-Order offers the option for Tubes and 384-well plates, is this option available for AmpliSeq HD?
The only order options available are, either Tubes Only or Plates Only option. If you need to order Tubes and Plates, we recommend placing 2 separate orders.
When I go through the Preview Order process for my AmpliSeq panels I always end up in the Order Preview page before I can proceed with the order. Has anything changed in the ordering process for AmpliSeq panels and which ordering options do I have?
The ordering process and the format/content of AmpliSeq panels remain unchanged.
The Order Preview page shown at the end of the preview order process includes both technical and ordering details meant to provide a detailed summary of the specific selections made before proceeding with the actual ordering of the panel (optionally, a list of additional recommended consumables based on the selected panel configuration can also be displayed and desired items can be added to the order as needed).
Once you are satisfied with the panel format configuration shown in the Order Preview page, depending on your account setup on thermofisher.com and your local/administrative regulations, you can either order your AmpliSeq panel directly online through the Thermo Fisher Cart (“Add to cart” button) or place an offline order (e.g., through a written purchase order) through your usual Customer Service channels based on the information provided on the same page. *
The Download SOS button at the bottom of the Order Preview page allows capturing and exporting in a text file (SOS Document) all the relevant information related to the chosen panel configuration. The SOS document is required when placing an order that is processed by Customer Service teams (e.g., purchase order sent over email).
In general, when attached to offline communications with Thermo Fisher Scientific personnel, the SOS Document can be used to facilitate the correct processing of your requests for the specific panel configuration of your interest (e.g., requesting quotes, raising technical support inquiries, etc.)
Orders placed in the Thermo Fisher Cart, do not require the SOS document.
Customer Service inquiries related to AmpliSeq orders can be sent by email to the following addresses (by region):
Americas: genomicorders@thermofisher.com
EMEA: uk.primers@thermofisher.com
Asia Pacific: orders.sg@thermofisher.com
Japan: jpprimerorder@invitrogen.com
China: ampliseq_cn@lifetech.com
*Different processes might be in place in countries served by distributors, which are also unchanged. Please refer to your usual local Thermo Fisher Scientific resources for more information.
I noticed a field called “Provided materials” in the Order Preview page, but I cannot find anymore indications on the number of reactions that can be processed with my Made-To-Order, Community or AmpliSeq HD panels. Is anything changed in the content or format for these panels and how do I know how many reactions I can process once I will receive my panel?
There has been no change in format, manufacturing process or materials delivered for any AmpliSeq panel, including Made-To-Order, Community and HD. The “Provided materials” field in the Order Preview page is meant to provide specific details on the delivered materials for the panel format chosen by the user (in addition to the general information for standard Made-To-Order and HD formats reported in this same FAQ section). Among other things, this information, combined with the AmpliSeq reaction setup details available in the pertinent User Guide (direct link also provided in the Order Preview page based on the panel format selection), is meant to be used to estimate the number of samples/reactions that can be processed given the selected panel format and library preparation method. Please contact your Thermo Fisher Technical Support resource in case help is needed in this respect.
I noticed that in the Order Preview page there is a “Chosen format” field in the Design configuration summary table for Made-To-Order, Community and AmpliSeq HD designs. What does the information reported in this field correspond to?
The information in the “Chosen Format” field is a text string meant to provide confirmation of the selection made by the user on the “format” button and on the “special instruction” menu during the order preview journey. For Made-To-Order and Community panels, the reported info is “format button, special instructions”; for AmpliSeq HD panels, only the “format button” info is reported (no special instructions menu available).
Note: For Made-To-Order panels, the “384-well plates only” special instructions menu entry is available under the “Tubes plus 384 well plates” format category button. Therefore, according to the above, the “Chosen Format” field for this selection is “Tubes plus 384 well plates, 384-well plates only”. As per the corresponding “Provided materials” field, only 384-well plates will be provided for this selection.
Why is a suffix sometimes added to the Design ID when ordering or requesting a web quote for certain Ion AmpliSeq™ custom panels, and does it affect the panel content or configuration?
The Design ID defines the content of an Ion AmpliSeq™ custom panel. For certain panel types – including Ion AmpliSeq™ Pre-Designed Panels, Ion AmpliSeq™ White Glove panels, and Oncomine™ tumor-specific Core panels – the same Design ID may be ordered by multiple users. In these cases, an additional alphanumeric suffix (preceded by “!”) may be appended during quote generation or order processing to create a unique transaction reference.
This suffix does not change the panel content or the selected order configuration. Panels that share the same Design ID correspond to the same underlying design, regardless of any appended suffix.
-
To submit human or mouse genomic targets for assay design submission, users can input a BED file of genomic regions of interest, or a Gene List file based on HUGO gene symbols and aliases.
-
The BED format files in AmpliSeq use the convention known as “zero-based, half-open” (ZBHO) coordinates, both for input and for output files. In contrast dbSNP and COSMIC use “one-based, inclusive” (OBI) coordinates. Notice then that compared to dbSNP and COSMIC, AmpliSeq coordinates will have a start coordinate one less than that shown on the dbSNP and COSMIC databases.
When comparing coordinates in BED files between AmpliSeq and data from the UCSC browser, please be aware that the UCSC Genome Browser uses both coordinate systems: OBI in the web interface and ZBHO in their database and data downloads.
What is the current turn around time for a submitted design with respect to the target size or the number of targets?
A design of 250kb or less should be returned in less than 48 hours of submission. For designs over 250kb or a large number of targets, you should expect a longer turn around time.
-
Yes, you can generate BED formatted files by utilizing the UCSC Genome Browser export feature in the Table Browser section.
Can we upload FASTA sequences?
Not at this time. We are currently exploring different methods for uploading regions to the Ion AmpliSeq Designer.
Can I use Galaxy instead of UCSC or IGV?
Yes. Any tools can be used to help you generate files for submission, but it is important to
make sure the correct version of the genome is being used (hg19 for human, mm10 for mouse).
What browsers are supported with this application?
We support Firefox, Google Chrome, Safari, and Internet Explorer 9 and above.
What can I do to ensure that an entire exon is covered in my design?
If coverage obtained from the initial design is less than 100%, you can try to extend the primer further out into the intron to capture the whole exon. Primer regions are not considered covered, so placing padding may ensure that we are able to get good quality sequence at the ends of exons, and to get some sequence read into the splice junction regions.
Is it possible to use the Ion AmpliSeq design to actually screen a large number of SNPs (up to a 1000 or more) in a large number of individuals (up to a 1000 or more)?
Yes, Ion AmpliSeq Designer allows you to do SNP genotyping by sequencing. Alternatively, you can also consider Taqman SNP Genotyping Assays for a large number of samples.
Is it possible to use the designer to detect differences between methylated/non-methylated DNA?
The Ion AmpliSeq Designer does not currently allow designing customized primers for methylation experiments. However, our White Glove team can assist creating a custom methylation panel for that purpose. Please contact your local representative for more info.
Can the designer be used for targeted whole genome sequencing?
The Ion AmpliSeq is used for targeted resequencing. It cannot be used to sequence whole genomes.
Exactly what is provided as output to the assay designs?
When you click on the Download Results button of your Results ready project/version, the
following output files are generated in a compressed folder.
| File Name |
Details |
|
#_coverage_summary.csv
|
Gene-specific and region-specific coverage details |
|
#_coverage_details.csv
|
This file provides details of coverage by exon for targets submitted by CDS or CDS+UTR (targets submitted as regions cannot be decomposed into exon-equivalents, so they are not listed in this file). If a request has no CDS (or CDS+UTR) targets, then there is no information for creating this coverage_details.csv file.
|
|
#_Submitted.bed
|
BED file with the genomic coordinates submitted to design |
|
#_Designed.bed
|
BED file of coordinates of what the application designed to |
|
#_Missed.bed
|
BED file of coordinates that were missed by the designer |
|
#_384WellPlateDataSheet.csv
|
This file provides the position of amplicons (or single primers) in the 384-well plates that are included with tubes and plates panel order formats (compatible panels only). For each amplicon or primer in the panel, the file reports the row and column coordinate of the corresponding well in the plate, along with info on reference, chromosome, amplicon insert start, and amplicon insert end. Empty plate wells have the value "Blank" in the "Amplicon_ID" column (and the other fields are left empty).
|
|
#_hotspot.bed
|
This file contains all hotspot variants that overlap with designed.bed coordinates (specific to each design) and an internally curated list of oncology hotspots. This file is available by default for AmpliSeq HD designs (IAH) and Oncomine Tumor Specific Panels (OAD if customized), and can be optionally generated for human custom DNA Made-To-Order designs.
|
|
#_amplicon_insert_size_histogram.png
|
This image file provides the user with a histogram view of the insert-size distribution of all the amplicons found in the panel. The insert being the region between the primers, which is amplified as reads, which in turn are used for sequencing of the targeted region.
|
|
#_amplicon_size_histogram.png
|
This image file provides the user with a histogram view of the amplicon size distribution of all the amplicons found in the panel. The amplicon being the targeted region including the primer information at the 5' end and 3' end.
|
|
plan.json
|
This file contains information to automatically configure a run plan for the panel, when the panel's files are directly downloaded from the Torrent Server 3.6
|
AmpliSeq primers manufactured by Thermo Fisher are optimized for their use with the Ion Torrent platforms. The primer sequences represent confidential information that cannot be disclosed to users or to any third party under any circumstance.
When I submit a UCSC .bed file with exons from a few genes the user interface estimation of the size of my design is very large. Why? how can I prevent that?
The user interface does not check for duplicate regions or any overlaps of the regions submitted in a .bed file. The UCSC .bed files typically contain duplicate regions for many quasi-identical transcripts. Too many overlapping regions may lead to a wrong estimate which may prevent the submission if the target size exceeds the currently allowed limit of 500 Kb.
A simple and effective way that may help to prevent this, is by running the UCSC .bed file through the program mergeBed from the BEDTools suite. This will create equivalent regions in a smaller .bed file.
Which is the largest design that I can submit to AmpliSeq?
The largest design that can be submitted directly to the pipeline is at most 500 Kb. However the pipeline is capable of processing designs up to 5 Mb, but such designs are predictably costly and take up a large number of computational resources.
In the cases of submissions larger than 500 Kb, the user will be contacted by email requiring more details about his/her interest in that particular, design and the design will be put on hold until the contact has been made.
What is different in AmpliSeq.com 7.2?
The DNA made-to-order pipeline in AmpliSeq 7.2 incorporates a number of algorithm improvements developed within the Custom Solutions Group at Thermo Fisher. These improvements have been validated on a large number of designs, and provide improved coverage and accuracy.
Will the changes in AmpliSeq.com 7.2 always improve coverage?
In the majority of cases, AmpliSeq.com will provide more coverage than previous versions. However, the new algorithm also implements additional checks and the quality of candidate oligos and amplicons, in particular, checking for specificity and insert uniqueness. In some less common scenarios, for example in cases of low-complexity regions of DNA, the algorithm may not be able to find amplicons meeting these quality checks to cover a region. In this situation, the reported coverage may be lower than previous versions, though actual amplicon performance should be improved.
-
AmpliSeq.com 7.2 will continue to generate multiple solutions corresponding to different stringency levels and numbers of pools. However, we have found that generating solutions with different target amplicon lengths were not needed in most cases. Consequently, AmpliSeq.com will now only generate solutions for the optimal amplicon length for the desired sample type. This only applies to AmpliSeq DNA made-to-order. See the next question for further details.
Which designs will be impacted by the changes?
In 7.2 the changes will affect AmpliSeq DNA made-to-order releases for human (hg19, GRChg38) and mouse (mm10) genomes. Improvements for additional genome support, including support for custom genomes, and support for AmpliSeq HD, are planned for a future release.
If I already created a draft design, would I still get all the solutions for all the amplicon sizes as I did before?
No. Once you're ready to submit your design for calculation, you will be prompted to select the desired amplicon size, and only solutions for the selected amplicon size will be generated.
When using the Copy Amplicons functionality sometimes I get extra pools or errors. Why does this happen and what can I do to get past the error?
When using the copy amplicons functionality, the system will attempt to create a solution which includes all the requested amplicons, if necessary adding additional pools, up to a maximum of 5 pools, in order to accommodate overlapping amplicons. If it is not possible to create a solution using the maximum number of pools then an error is raised. In this case the user must remove some overlapping amplicons/targets in order to create a valid solution.
What is the maximum number of characters that I can use to describe a region name and which characters are not supported
Region names can be at most 70 characters long, and may not contain any of the characters '/', '"', ''', '=', ',', ';', '[', ']', '\', '|', space or tab.
Are there any restrictions that I should be aware of when uploading target bed files?
Uploaded target bed files must have the extension '.bed' (lower case). There are no other restrictions.
On occasion, there are gene symbols that are not recognized by the system, what can I do in this case?
If a gene-symbol is not currently in the known list of aliases then it will be necessary to upload the specific regions included in the gene, in order to design a solution for the gene.
For AmpliSeq Custom Made-to-Order, can I copy an amplicon from an FFPE design to a cfDNA design?
Yes, please refer to the following compatibility table to understand the changes done during the Copy amplicons event:
| Source DNA Type |
Initial Draft DNA Type |
New Modified Draft DNA Type |
| cfDNA (140bp) |
cfDNA (140bp) |
No modification |
| cfDNA (140bp) |
FFPE (175bp) or Standard (275 or 375bp) |
No modification |
| FFPE (175bp) |
cfDNA (140bp) |
FFPE (175bp) |
| FFPE (175bp) |
FFPE (175bp) or Standard (275 or 375bp) |
No modification |
| Standard (275bp) |
Standard (275bp or 375bp) |
No modification |
| Standard (275bp) |
cfDNA (140bp) or FFPE (175bp) |
Standard (275bp) |
| Standard (375bp) |
Standard (375bp) |
No modification |
| Standard (375bp) |
cfDNA (140bp) or FFPE (175bp) or Standard (275bp) |
Standard (375bp) |
For AmpliSeq HD Custom Made-to-Order, can I copy an amplicon from an FFPE design to a cfDNA design?
Yes, please refer to the following compatibility table to understand the changes done during the Copy amplicons event for AmpliSeq HD:
| Source DNA Type |
Initial Draft DNA Type |
New Modified Draft DNA Type |
| cfDNA (125bp) |
cfDNA (125bp) |
No modification |
| cfDNA (125bp) |
FFPE (175bp) |
No modification |
| FFPE (175bp) |
cfDNA (125bp) |
FFPE (175bp) |
| FFPE (175bp) |
FFPE (175bp) |
No modification |
Can I copy amplicons from an AmpliSeq Custom Made-to-Order design to an AmpliSeq HD Custom Made-to-Order draft design?
Yes, please refer to the following compatibility table to understand the changes done during the Copy amplicons event where the draft design is for AmpliSeq HD:
| Source DNA Type |
Initial AmpliSeq HD Draft DNA Type |
New Modified Draft DNA Type |
| cfDNA (140bp) |
cfDNA (125bp) |
FFPE (175bp) |
| cfDNA (140bp) |
FFPE (175bp) |
No modification |
| FFPE (175bp) |
cfDNA (125bp) |
FFPE (175bp) |
| FFPE (175bp) |
FFPE (175bp) |
No modification |
| Standard (275 or 375bp) |
cfDNA (125bp) |
Only amplicons <200bp will be copied, others will be ignored. Draft will be converted to FFPE type (175bp) to accommodate as many amplicons as possible. |
| Standard (275 or 375bp) |
FFPE (175bp) |
Only amplicons <200bp will be copied, others will be ignored. Draft will remain as FFPE type (175bp). |
Can I copy amplicons from an AmpliSeq HD Custom Made-to-Order design to a regular AmpliSeq Custom Made-to-Order draft design?
No, it is not possible to copy amplicons from an AmpliSeq HD design to a regular AmpliSeq design.
-
Coordinates in the Polymorphism.bed file indicate regions of the sequences in the custom reference FASTA file with high polymorphism (i.e., SNPs, indels, or other variation). Ion AmpliSeq Designer avoids overlapping primers with such regions since performance may be unpredictable. Consequently, in regions with many polymorphisms, Ion AmpliSeq Designer may fail to find suitable primer locations in order to sequence these regions. In this situation it may be necessary to resubmit the Polymorphism.bed file with very rare polymorphisms (e.g. minor allele frequency < 1%) removed.
When I use the Copy Amplicons functionality, why do the amplicon pool assignments often change in the new design?
The Copy Amplicons function ensures that all amplicons copied from a source design are included in the new design. The assignment of amplicons to pools is re-generated for each design with the aim of balancing the pools and avoiding any inter-amplicon conflicts. Balancing of the pools ensures that you have a similar number of amplicons in each pool. As a result, whenever amplicons are added to, or removed from a design, the pool assignments of other amplicons within the design is likely to change.
Does having a hotspot.bed file in my panel results mean that only those variants can be detected?
No. A hotspot BED file defines region targets that typically represent relevant variants. Specifying a hotspots file to use in a run is optional and enables Torrent Variant Caller to identify and report if a specific variant is either present, absent or could not be determined in the positions defined in the file. Specifically, a hotspot file instructs the Torrent Variant Caller to always include these variants in its output files, including evidence for called variants and filtering threshold(s) that disqualified a variant candidate in case a call could not be made. A hotspot file affects only the variantCaller plugin, not other parts of the analysis pipeline. If you don't specify a hotspots file, the software output includes only the variants detected between your sequence and the reference genome.
Why did I get a warning banner regarding the automatic CNV design functionality being ignored when submitting a design with gene targets flagged for CNV that also contained copied amplicons from previous design(s) (partial subset design)? How can I combine CNV-compatible gene targets with existing amplicons from previous designs?
Copied amplicons are considered must-have content for a subset design, therefore they will always be included in the final design. In case of conflict with additional newly designed amplicons (including amplicons for CNV gene targets), the copied amplicons will always prevail (i.e., they will be those retained in the final design at the expense of the new amplicons). This could potentially impair the automatic CNV design pipeline in unpredictable ways. Therefore, when a Made-To-Order design is submitted, if copied amplicons are detected alongside with CNV gene targets, then the automatic CNV design functionality is switched off and the warning banner is presented to the user for confirmation of the same.
For the same reason, combining CNV-compatible gene targets with existing amplicons is currently not supported.
I am interested in gene CNV targets on sex chromosomes. Can I use the automated gene level CNV design functionality to add them to my design? Is there any specific requirement I should consider?
It’s not possible to detect CNVs on sex chromosomes if all the targets on a panel are on the sex chromosomes (additional amplicons must also be added to autosomal chromosomes by adding targets accordingly). The automatic gene level CNV design functionality can be used to add CNV gene target on sex chromosome, but it does not automatically check for the presence of additional autosomal targets. Therefore panel-level CNV compatibility when sex chromosome CNV targets are present must be manually verified by the user.
How does the software accommodate intronic regions?
When the user submits a gene to design, only exons are used as targets. If you wish to design
across the whole gene (exons and introns) the user needs to submit the start and end coordinates
of the gene.
-
No. The designer uses exon coordinates as listed by the UCSC Genome Browser. Promoters are not
part of the exons and need to be requested using a BED file describing the genome coordinates.
What is the level of overlap among the primers? Are the overlapping primers in the same tube?
Primers in the same tube do not overlap. As our product line evolves this might change in the
future and a small overlap might be possible.
For the Ion AmpliSeq Designer, are primer sets designed automatically (with a computer program), without interrogation from a research scientist?
The process is an automated pipeline, optimized to provide the maximum coverage with reliable primer sets.
How are the Ion AmpliSeq Custom designs validated?
Each primer pool goes through a rigorous process to meet strict design specifications. During
the design of our pipeline, we validated a substantial number of our custom assays though wet
lab testing.
If I submit two continuous regions (175 - 225 bp range each) combined as one BED file, is it possible to get the designed primers for the overlapping region?
If an overlapping region is submitted to the design pipeline, internally the region is concatenated and treated as a single region for design, thus there will be no overlap. The two regions are reported back in the UI as submitted. While it is possible that an amplicon might be prorated twice, once in each of the original regions, this amplicon (and its primers) only occurs once in the design (see the plate file).
What is a superamplicon?
A superamplicon is created when two forward PCRs joined to form one large amplicon. The pooler algorithm in the pipeline separates primers into separate pools to minimize this.
The BED file specifications state that in a BED file the chrStart number is zero-indexed and the chrEnd number is not included in the feature. Are you following this convention for upload and are the numbers shown in the designer 1-indexed or 0-indexed?
chromStart - The starting position of the feature in the chromosome or scaffold. The first base in a chromosome is numbered 0.
chromEnd - The ending position of the feature in the chromosome or scaffold. The chromEnd base is not included in the display of the feature. For example, the first 100 bases of a chromosome are defined as chromStart=0, chromEnd=100, and span the bases numbered 0-99.
Can you describe more about how the Ultraplex technology works?
Development work from over a decade allows us to produce primer designs that allow simultaneous amplification of many amplicon targets. A unique chemistry has been developed for Ion AmpliSeq that allows removal of any primer dimer formed along with the majority of the primer itself from the amplified template. This makes sequencing very efficient by not wasting bases on non-informative primer sequence and allows for very clean sequencing reactions.
Do your designs take into account the presence of pseudogenes?
Yes. The pipeline first attempts to design primers that only match the target, and not the pseudogene (or duplicate) version(s). If the target gene is not covered in the initial rounds of primer selection, then the match parameters are relaxed, for the sake of coverage, in later rounds, attempting to maintain the uniqueness of the inserts.
If two amplicons overlap, do the primers produce a big product in addition to two small ones?
The pooling step in the design is optimized in order to minimize the interference between overlapping amplicons. Hence, overlapping amplicons would be segregated into different pools.
Why my gene is not accepted for design?
There are several reasons that explain why this happens:
- A gene must be part of the UCSC Reference Gene dataset
- A gene must have at least one coding transcript
- A gene must not map to more than one genomic location (this includes pseudoautosomal genes (PAR1,2) )
- A gene must not map to un-assembled contigs or alternate assemblies - examples for human include: chrUn_gl000228, chr4_gl000194_random and chr6_cox_hap2 (see the UCSC FAQ on chrN_random tables )
Which sequence versions does AmpliSeq use in its computations for panel designs?
DNA
Gene targets correspond to RefGene v98 for Human (hg19) and v99 for Human (hg38)
| Genome |
Genome version |
dbSNP |
COSMIC |
| Human (hg19) |
Feb. 2009 (GRCh37) |
v156 |
v97 |
| Human (hg38) |
Dec. 2013 (GRCh38.p2) |
v156 |
v97 |
| Mouse (mm10) |
Dec. 2011 (GRCm38) |
v150 |
N/A |
RNA
Human RNA Canonical RefSeq Transcripts* - Feb. 2009 (hg19, GRCh37)
HGNC Database, HUGO Gene Nomenclature Committee (HGNC), EMBL Outstation - Hinxton, European Bioinformatics Institute, Wellcome Trust Genome Campus, Hinxton, Cambridgeshire, CB10 1SD, UK http://www.genenames.org, 11/2012
*For compatibility reasons, Spike-in designs use the RefGene version of their corresponding pre-designed AmpliSeq On-Demand panel (v74).
How does hotspot target prioritization work and what is it meant for?
When non-overlapping amplicons covering adjacent submitted hotspot targets are not available for a one-pool design (example figure below), the hotspot pipeline will compare overlapping amplicons ("a" and "b", in the example) and eliminate one or more of them, resulting in the missed coverage of the targets they covered.
The target prioritization functionality allows the user to define a priori which hotspot target(s) the design pipeline should attempt to retain when such conflicts by overlapping amplicons occur. If hotspot targets are prioritized by the user, then the design pipeline will attempt to resolve conflicts by preferentially retaining amplicons, if available, that cover the prioritized targets, at the expense of non-prioritized ones. Although under certain circumstances target prioritization might also resolve conflicts in 2-pool designs, this functionality is mostly relevant for 1-pool hotspot designs where overlapping amplicons are mutually exclusive (i.e., cannot be all retained in the final 1-pool design).
Note: The hotspot target prioritization functionality is not meant to be used for increasing the overall coverage of 1-pool hotspot designs (in case, lower specificity solutions can be used for the scope, see FAQ section), but instead, if suitable amplicons are available for the scope, to try and prioritize the coverage of hotspot targets most relevant for the user when overlapping amplicon conflicts occur due to target proximity.
|
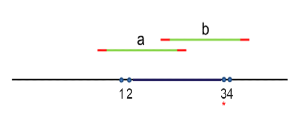
Effect of prioritization: With reference to the above figure, if target 3* is prioritized by the user at the hotspot design submission step, then the design pipeline will retain amplicon "b" for the 1-pool design solution and the prioritized hotspot target 3 will be covered (along with target 4). Amplicon "a" will instead be eliminated resulting in the missed coverage of hotspot targets 1 and 2.
|
How do I transfer a design that I created from a Custom Reference to a new account in AmpliSeq.com?
Please contact your our Global Support team (ampliseq-designs@thermofisher.com) to request the transfer. Note that the transfer will be done not just for the requested design, but also for all designs associated with the same Custom Reference. This is an all or nothing event.
I requested the transfer of a design created using a Custom Reference to a new account, however, I am unable to find other designs that I had created using the same Custom Reference, where did they go?
When transferring a design created from a Custom Reference, all designs must be transferred along with the reference. The other designs should now be found in the recipient account.
How do I share a design that I created using a Custom Reference?
a) You can share designs whether they are from Reference Genomes like (hg19, mm10 and others), as well as designs created from Custom References using the “Sharing” functionality found on the main design page, next to the Download results button.
b) After selecting the “Sharing” button, you will need to enter the full email address of the recipient you wish to share the design to. Please note that only registered emails will be recognized as valid recipient email address. Make sure that the recipient does have a valid AmpliSeq.com account.
What are the limitations of shared designs?
a) Share designs are meant to be used for review, analysis and ordering purposes. You may not create new designs from a shared design. For this reason, the “Copy Amplicons” and “Copy Targets” functionality is disabled.
b) The sharing of a design can only be done by the original creator of the design. A recipient of a shared design, cannot re-share the same design to another user.
What is the implication of supporting NCBI genomic references?
We have decided to adopt NCBI formatted references as our standard for uploading Custom References to AmpliSeq.com. This means that users will be able to download references files from the NCBI databases.
-
Please visit the help section for Custom References Working with Custom References for further information about all the requirements needed for successfully upload your Custom Reference.
Is there a way to edit the Known polymorphism (BED) file without having to re-upload the Custom Reference?
Yes, in our latest release we now allow the removal and re-upload of the Polymorphism (BED) file, without the need for a new Custom Reference.
Is there a way to download/access a Polymorphism (BED) file that has been removed from an existing Custom Reference?
No, only the latest Polymorphism (BED) file associated with the Custom Reference is available from the Design References page. If the file is removed, it can no longer be accessed or retrieved.
Is hg38 the same genome version as GRCh38?
Yes, they are the same version of the human genome. GRCh38 stands for “Genome Reference Consortium Human Reference 38” and it is the primary genome assembly in GenBank; hg38 is the ID used for GRCh38 in the context of the UCSC Genome Browser.
Essentially, how is hg38 different from hg19?
The hg19 build is a single representation of multiple genomes. The hg38 build provides alternate sequences (“alt_sequences”) for some genomic regions for which their variability prevents adequate representation by one single reference.
How is the hg38 reference used by Thermo Fisher Scientific software different from the references publically available from places like UCSC or NCBI?
• The version used by our software is based on GRCh38.p2 (http://www.ncbi.nlm.nih.gov/assembly/GCF_000001405.28)
• Unplaced, Alternate and Unlocalized contigs are listed as separate chromosomes and ordered first by chromosome localization, then by the alphabetic order of the Genbank accession of the contigs.
• Repeat and SNP locations are soft-masked into lower case letters, while the ambiguous IUPAC bases, duplicated centromeric arrays, and chrY PAR regions are hard masked into 'N's.
• It contains chr1-22, chrX, chrY, and chr22_KI270879v1_alt.
• Contig chr22_KI270879v1_alt is hard masked except region 269814-279356 (1-based).
• Gene GSTT1 is located at chr22_KI270879v1_alt:270308-278486.
I know there are many more “alt_sequences”, why is that your version of hg38 only considers one of those?
Our version of hg38 only considers the chr22_KI270879v1_alt. This alt chromosome contains gene GSTT1 that was part of chr22 in hg19. This was an internal decision which was made to enable standardization of the genome reference for use across multiple businesses within our organization.
-
We strongly recommend that you download our version of the hg38 from our website. This version is the one that is assumed in all of our software applications and has been tested for compatibility.
-
At this moment we do not offer any conversion tools. We recommend our users touch base with their own bioinformatics experts for further guidance.
Can I analyze old assays or panels with the new hg38 build?
No. The old assays and panels were created using hg19 as a reference and should be analyzed with the tools and analysis pipelines created for hg19.
Can I still design, order and analyze old AmpliSeq panels based on hg19?
Yes. The pipelines and tools for using hg19 as reference for design and analysis are still available.
Can I still annotate my old variants (based on hg19) with Ion Reporter Software?
Yes. The variant calling workflow based on hg19 will be available in Ion Reporter. If your design was created using hg38, then you can also call and annotate variants using Ion Reporter.
Can I copy amplicons from an hg19 design to an hg38 design (or vice versa)?
No. Amplicons from a custom design can only be copied to another custom design associated with the same reference. It is not possible to copy amplicons to a custom design associated with a different reference even if both references are human.
Will there be a new version of the Kitted and Community AmpliSeq Research Panels based on hg38?
Not at this moment. The off-the-shelf panels (ie exome), On-Demand panels and Community panels will still be based on hg19. Conversion of pre-designed panels may be considered in the future based on market demand.
Will there be a version of the Oncomine Panels based on hg38?
Not at this moment. The Oncomine panels will still be based on hg19.
Are your Ion Reporter Software annotations based on hg38 or hg19?
If your AmpliSeq design has been created using hg38 as a reference, then you can create an ad-hoc workflow in Ion Reporter for analysis. All analysis and annotations will take in consideration hg38 as a reference. However, at this moment there are no hg38 workflows in Ion Reporter. The tools for analysis and the annotations for hg19 will still be available.
Why does the Ion AmpliSeq RNA pipeline not accept the entered Gene Symbol?
The pipeline recognizes HGNC approved gene symbols, previous gene symbols, and synonyms. It is case sensitive. Please check the Gene Symbol entered for typos and then search for the Gene Symbol on the HGNC website , to confirm that is a valid entry. The pipeline requires that the Gene Symbol entered is unambiguous, meaning that it resolves to a single HGNC approved gene symbol.
Why does the Ion AmpliSeq RNA pipeline not produce designs to comprehensively target all RefSeq transcripts of the entered Gene Symbol?
The pipeline uses strict criteria in order to design a single assay per gene that will result in consistent amplification while minimizing the risk of amplifying genomic DNA, this is done by targeting splice sites between exons. If there is not a splice site that is shared between all RefSeq transcripts of the entered Gene Symbol or if a passing assay could not be designed to the most common splice site then an assay will be produced that is compatible with a subset of the RefSeq transcripts for the entered Gene Symbol.
Why does the Ion AmpliSeq RNA pipeline not accept the entered RefSeq accession?
The pipeline attempts to resolve a single hg19 genomic alignment, where an exception is made for pseudoautosomal alignments, for each RefSeq accession. Unrecognized or Invalid RefSeq transcript accessions can result when the entered accession has been permanently suppressed or was not successfully resolved to a single hg19 genomic alignment.
Why does the Ion AmpliSeq RNA pipeline produce a design that is not specific to the entered RefSeq transcript accession?
The pipeline will design a single most-inclusive assay for the gene that corresponds to the entered RefSeq transcript accession. To achieve this, the pipeline will attempt to design an assay to the most common splice site found in the RefSeq accessions for the gene.
-
No. These are among the list of features that are on the development roadmap and will be included in subsequent releases of the Ion AmpliSeq RNA pipeline.
Is there a way in the AmpliSeq RNA pipeline to specify genomic regions to design against?
No. The RNA pipeline does not allow specifying genomic regions for design.
-
No. The RNA pipeline does not allow the entry of FASTA sequences for design.
Why the AmpliSeq RNA pipeline designed.BED file isn’t compatible with UCSC genome browser?
The data on which AmpliSeq RNA pipeline is based on, is the NCBI's RefSeq sequences. A consequence of this is that all coordinates in the .BED files correspond to RefSeq coordinates that are not recognized by the UCSC genome browser. However the .BED files produced by the RNA pipeline are fully compatible with other pieces of IonTorrent software.
Why is there a restriction on the minimum and the maximum number of amplicons in my design?
The minimum of 12 amplicons is due to chemistry performance limitations with low number of amplicons per pool. The exception to this limitation are spike-in designs which allows < 12 amplicons/pool, since these panels are exclusively meant to be used to physically add amplicons to larger main designs to expand their content.
Note: if an MTO DNA design panel has fewer than 48 amplicons it is subjected to our minimum order quantity policy of 48 amplicons (96 oligos) and its associated pricing.
The maximum number of amplicons in RNA designs is based on validation test results that demonstrated good gene expression dynamic range on a sequencing run. A good dynamic range means good sensitivity to detect extremes in the expression of genes in a given sample.
How can I find out which is the TaqMan assay (if any) corresponding to my RNA targets?
The last two columns of the IAD#_DataSheet.csv file (included in the download results files for an RNA design) reports the preferred TaqMan assay that can be used for verification of the particular target and classification of the TaqMan assay with 2 possible values:
• Classification 1: The recommended TaqMan Gene Expression Assay targets the same exon or exon-exon boundary as the Ion RNA AmpliSeq Design.
• Classification 2: The recommended TaqMan Gene Expression Assay targets the same set of RefSeq Accessions as the Ion RNA AmpliSeq Design.
Follow this link for instructions on how to get the corresponding TaqMan assay.
AmpliSeq RNA warns me about including high expressed genes in my design. How do you know they are high expressers?
The high expressed genes were selected from rank-ordered lists of whole transcriptome RNA-Seq expression data derived from universal human reference RNA (UHRR, Stratagene). UHRR is comprised of purified RNAs from 10 distinct human cell lines and has been shown to be an accurate and reproducible standard for comparison of gene expression data. This reference RNA has been utilized by the highly referenced Microarray Quality Control (MAQC) consortium as well as the more recent Sequence Quality Control (SEQC) study. There is familiarity among microarrays users primarily as well as some RNA-Seq customers around the MAQC samples which also bolsters the decision to use this RNA for the rank ordered list.
Data Collection: Whole transcriptome sequence data was collected from both PGM and Proton runs using Ion 318 and Ion P1 chips, respectively.
Since UHRR represents several different tissues and we have a wealth of internal RNA-Seq data from this sample, we found UHRR to be a reasonable sample type for determining a common list of highly expressed genes.
References
UHRR product information
MAQC Study Webpage (FDA)
MAQC Paper
If one of the important genes missed in the coverage or was off the target, do we need to start all over? Could we just redesign that gene and plex it again?
No, you do not need to start all over. You can add, delete and edit genes or regions from the design results. You can even create a new version of the same design, with iterative modifications, and continue to resubmit the designs.
Is the recommendation to validate the sequencing data with Taqman SNP? (Direct sequencing vs indirect TaqMan assay?)
Variant confirmation can be done with other platforms, including Taqman SNP Genotyping Assays and Sanger sequencing–capillary sequencing.
Suppose we are targeting a region and the AmpliSeq Designer suggest a design consisting of 2 primer pools. For each sample, should we prepare a library for each amplification (each pool) or should we combine the 2 amplifications (the products of the 2 amplified pools), and then do the library?
If your design results in multiple pools and you are preparing libraries manually, then as per User Guide each pool should be amplified independently and amplification product shall be combined before primer digestion step for each sample.
Could the tumor-amplified DNA and normal-amplified DNA be loaded onto the same chip, then the sequences be separated out during analysis?
Yes, you can pool your tumor and normal samples together into a single chip run (assuming that your target size and required coverage can be achieved in a single chip). Many people perform differential pooling so that the coverage of the normal sample is lower than the tumor sample.
We see a large variation in the coverage of different regions of our panel. What could you recommend?
We recommend using the Ion AmpliSeq Library Plus Kit as it is expected to produce higher library yield, increased uniformity and result in more robust library amplification than the standard Ion AmpliSeq 2.0 Library Kit.
Is there any way to determine if a variation is true or is a mistake introduced by the polymerase?
There are a number of ways to perform orthogonal validation of mutations found by NGS, including TaqMan Genotyping Assays in standard or digital PCR formats, TaqMan Mutation Detection Assays for specific somatic mutations, or Sanger Sequencing using Capillary Electrophoresis.
-
To ensure that an entire exon is covered, by default we add 5 bp of padding up and down-stream of the selected target region to allow room to place the primers. Padding ensures that we are able to get good quality sequence at the ends of the exons and to get some sequence read into the splice junction regions. Primer regions are not considered covered. Therefore, if coverage obtained from the initial design is less than 100%, we can try once more to extend the primer further out into the intron to capture the whole exon.
Could you target multiple pathogens, while filtering out off-target human genome amplification?
Designs for pathogens are not currently supported for Ion AmpliSeq Designer.
Could I do two or three different amplifications and then pool before going into library prep?
It is possible to run 3 different AmpliSeq designs each with barcodes and combine them going into Template Prep.
Can I import pre-design PCR primer sets and validate if they work?
Target regions from pre-designed PCR primer sets can certainly be imported and submitted for design into the Ion AmpliSeq Designer. The specific parameters of Ion AmpliSeq Designer may result in primer sets that are different from initially pre-designed PCR primer sets but are optimized for use with the Ion AmpliSeq Technology.
Why am I getting fewer targets in my results than what I submitted in my .BED file?
Due to quality control considerations after submission, amongst other properties, the .BED file is reviewed for duplicates. Once the duplicates have been removed and other filters have been passed, the (probably reduced) file is accepted into the pipeline.
Why is my design marked as "Undesignable"?
The most common cause of a design request being marked as "Undesignable" is that AmpliSeq Designer is unable to generate any amplicons overlapping the target regions which satisfy the minimum design requirements.
Possible reasons for this include repeated sequences (e.g. pseudo-genes); low-complexity regions; very high or very low GC content or other problem sequences. These factors may prevent AmpliSeq Designer from being able to find pairs of primer locations which meet the minimum criteria for specificity, primer melting temperature, GC content, and so on. In such cases, we recommend checking the target sequences using a tool such as the UCSC genome browser. When using a custom reference, it is recommended that the sequence is manually checked by the user for the presence of duplicated or other problematic regions that might cause design requests to be undesignable. Note that, when uploading a custom reference, if the user selects an associated organism for specificity checks, the associated organism reference genome will be used in searching for repeats.
In addition, when using the Copy Amplicons functionality, AmpliSeq Designer will mark a design request as "Undesignable" if it is not able to fit all the copied amplicons into the available pools due to amplicon conflicts. Note that, while for AmpliSeq Custom designs, two amplicons conflict only if they overlap by more than the maximum allowable number of bases, in AmpliSeq HD some non-overlapping amplicons may also be flagged as conflicting due to other limitations of the AmpliSeq HD chemistry. In this situation, it may be necessary to remove some of the copied amplicons from the design.








